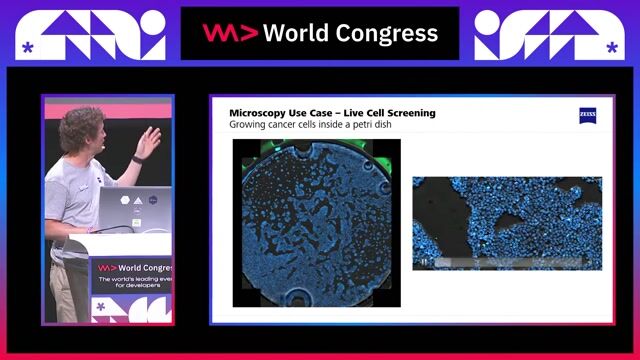
Stage: AI-powered multimodal integration of spatial omics data
Centre de Bioinformatique de Bordeaux
Canton of Bordeaux-2, France
19 days ago
Role details
Contract type
Internship / Graduate position Employment type
Full-time (> 32 hours) Working hours
Regular working hours Languages
EnglishJob location
Canton of Bordeaux-2, France
Tech stack
API
Artificial Intelligence
Artificial Neural Networks
Bioinformatics
Cluster Analysis
Software Documentation
Computational Biology
Python
Machine Learning
Open Source Technology
TensorFlow
PyTorch
Large Language Models
Deep Learning
Information Technology
HuggingFace
Integration Frameworks
Job description
- State-of-the-art Benchmarking: Evaluate existing multi-modal integration frameworks (e.g., SpatialGlue, GLUE, or Seurat v4) on public and in-house paired ST+MSI datasets to identify specific failure points in metabolic signal alignment.
- Methodological Development: Design and implement a novel Graph Neural Network (GNN) architecture to bridge the identified gaps.
- Implementation of Joint Embeddings: Develop an automatic supervised graph-learning workflow to map both modalities into a shared latent space, ensuring robustness against differences in spatial resolution and technical noise.
- Validation: Apply the developed model to in-house PDX glioblastoma datasets to characterize ROIs and validate the biological relevance of the integrated signal in collaboration with our cancer biology partners.
- Software Documentation: Deliver a clean, well-documented Python codebase (compatible with the lab's existing DIMet/SpacePath ecosystem)., * Galvis J, Guyon J, Dartigues B, Hecht H, Grüning B, Specque F, Soueidan H, Karkar S, Daubon T, Nikolski M. DIMet: An Open-Source Tool for Differential Analysis of Targeted Isotope-Labeled Metabolomics Data. Bioinformatics 2024 40 (5) btae282. https://doi.org/10.1093/bioinformatics/btae282.
- Galvis J, Guyon J, Daubon T, Nikolski M. Using DIMet for Differential Analysis of Labeled Metabolomics Data: A Step-by-Step Guide Showcasing the Glioblastoma Metabolism. Bio-Protoc. 2025 15 (2) e5168. https://doi.org/10.21769/BioProtoc.5168.
- Ravi VM, Will P, Kueckelhaus J,… & Heiland DH Spatially Resolved Multi-Omics Deciphers Bidirectional Tumor-Host Interdependence in Glioblastoma. Cancer Cell 2022 40 (6) 639-655.e13. https://doi.org/10.1016/j.ccell.2022.05.009.
- Wheeler K, Gosmanov C, Jimenez Sandoval M, Yang Z, McCall LI. Frontiers in Mass Spectrometry-Based Spatial Metabolomics: Current Applications and Challenges in the Context of Biomedical Research. TrAC Trends Anal. Chem. 2024 175 117713. https://doi.org/10.1016/j.trac.2024.117713.
- Liu T, Fang Z.-Y, Zhang Z, Yu Y, Li M, Yin M.-Z. A Comprehensive Overview of Graph Neural Network-Based Approaches to Clustering for Spatial Transcriptomics. Comput. Struct. Biotechnol. J. 2024 23 106-128. https://doi.org/10.1016/j.csbj.2023.11.055.
Requirements
- M2 / engineering school student in Computer Science, Applied Mathematics, AI, bioinformatics.
- Solid Python skills, including familiarity with deep learning frameworks (PyTorch, TensorFlow) and/or LLM APIs (Huggingface, OpenAI).
- Knowledge of machine learning.
- Prior exposure to omics data is a plus but not necessary.
- Background in molecular/cellular biology or metabolism is a plus; motivation to work in an interdisciplinary setting is essential.
The internship will provide hands-on experience at the intersection of AI and computational biology.
About the company
Despite recent advances in multi-modal spatial omics integration, metabolic semantics are still poorly used by existing frameworks: the key molecular modality (MSI-based spatial metabolomics/lipidomics) differs from ST in resolution, quantitativeness and section-to-section registration [4]. Most ST-MSI integration across adjacent sections relies on geometric registration (histology-derived anchors), however, invasive cells are sparse and white-matter anatomy is anisotropic, which can lead to biological signal misalignment. On the other hand, graph-based fusion methods (e.g., SpatialGlue, SpatialMET, COSMOS) improve integration in more homogeneous settings.