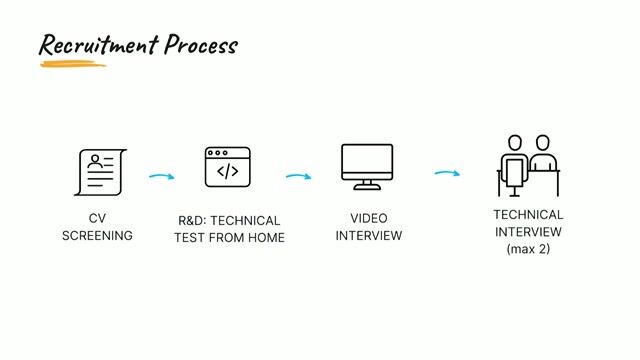
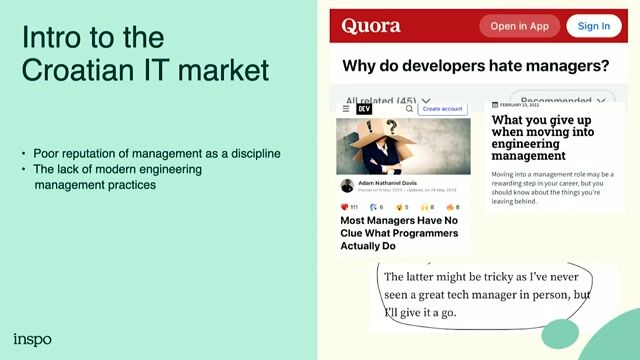
Job offer
Role details
Job location
Tech stack
Requirements
Master Degree or equivalent
Skills/Qualifications
§you possess a master degree in computational chemistry, bioinformatics, or a related field
§you have mastered molecular modeling techniques, machine learning algorithms, and programming languages like Python
§you are highly collaborative, with excellent communication skills and a passion for interdisciplinary research
§you like tackling complex problems, working at the interface of biology, chemistry, and computer science, and publishing your results in top-tier journals Internal Application form(s) needed 2026-ICN06 English version.pdf English (489.13 KB - PDF)